Thanks to rhi for providing code for complementing without reversing. Since my R programming skills are “limited”, comments and suggestions are welcome! However, the versatility of R allows you to automatically retrieve the reverse complement and (for example) save each of the primer in a different text file.Īlso, there is a nice library in R (seqinr) which can reverse complement and perform several other tasks ( ). Only now I found a web site in which you can copy and paste the primers on different lines and get the reverse complement of each primer on a different lines. The advantage of using this piece of code is that it is possible to automatically reverse complement a series of sequences: I had several primers to reverse/complement and I didn’t want to copy and paste them every time.
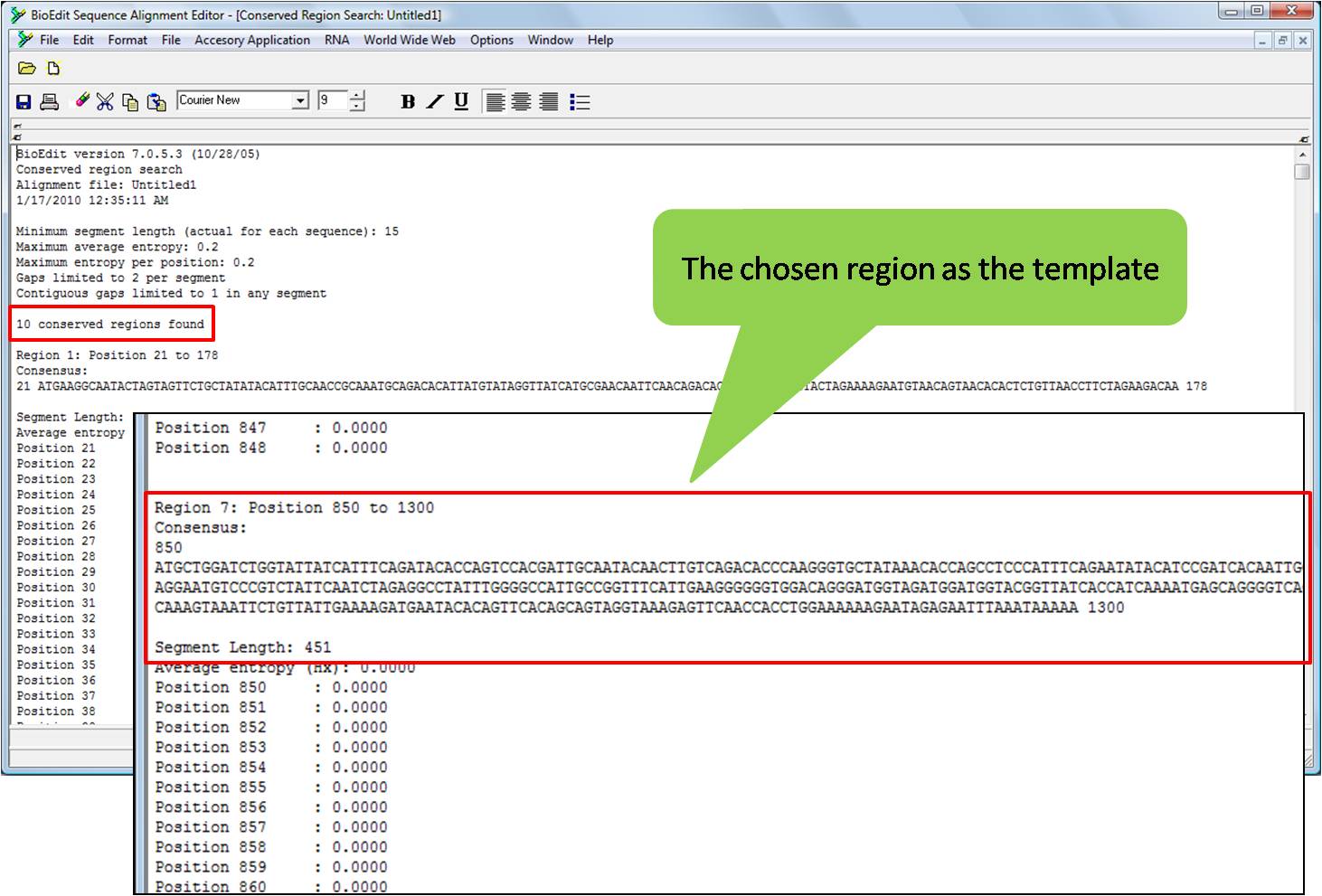
There are several web sites which can easily complement and reverse a DNA sequence (and RNA as well). The complemented (and eventually reverse) sequence, as a character vector.
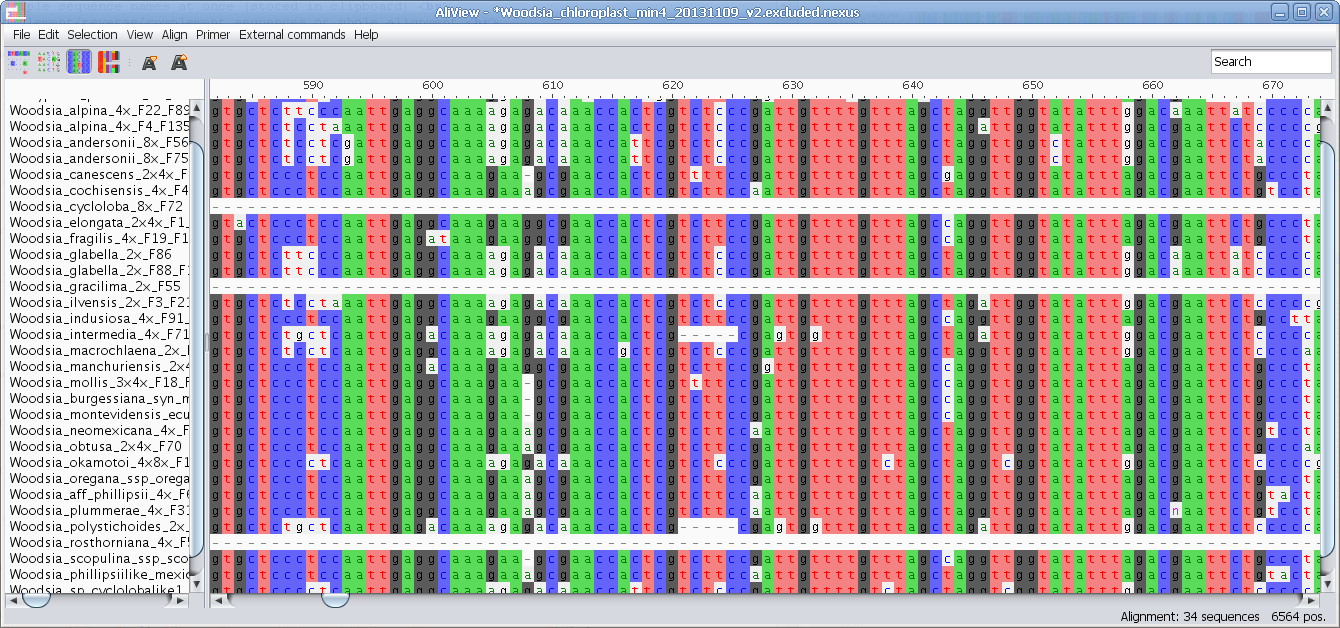
If TRUE, the function will return the reverse complemente, if FALSE, it will return the complementary sequence. You can complement and reverse complement, but not just reverse.Author Fabio Marroni ( ) Using a combination of the two you can reverse, complement, and reverse complement sequences as well.Ĭomplements (and eventually reverse) a DNA sequence, which has to be inserted as a character vector, no matter if lower or uppercase.Ģ) Cannot reverse without complementing.
#Bioedit reverse complement manual#
For difficult alignments that require a lot of manual gap adjustment, CodonCode Aligner supports round trip editing: export your aligned sequences, edit in you favorite editor like Se-Al or BioEdit. You can edit your contig alignment right in CodonCode Aligner.
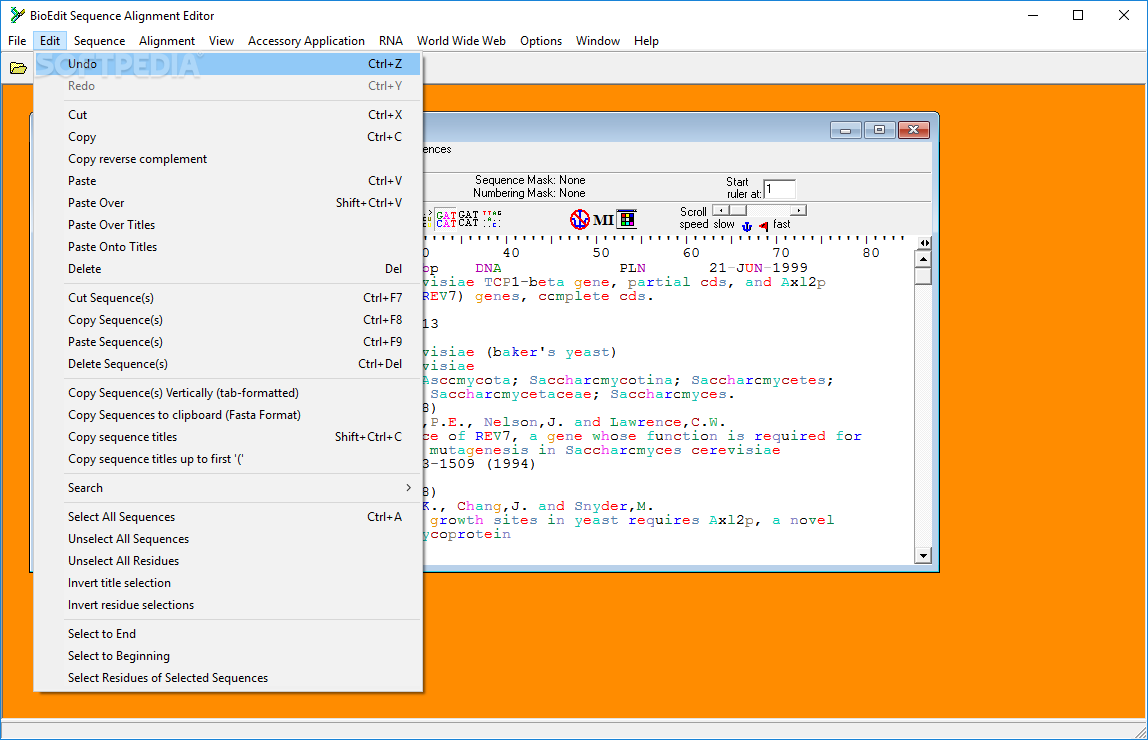
I would like to remind everyone (me in first place!) that the comp() function of the ( seqinr) package can complement a DNA sequence, and rev() function of Rbase can reverse a character vector. If necessary, CodonCode Aligner will reverse-complement contigs before alignment. This post is intended for documentation only.
